YU/BIU Summer 2022 STEM Research Showcase
YU/BIU Summer 2022

Washington University in St. Louis
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Hagay Shpaisman and of PhD student Udi Greenberg.
This abstract was previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book .
Title: Three-dimensional nanoparticle assembly by a modulated laser-induced microbubble for fabrication of a micrometric pattern.
Topic: Engineering, nanotechnology, directed materials assembly.
ABSTRACT:
Three-dimensional nanoparticle assembly by a modulated laser-induced microbubble for fabrication of a micrometric pattern
Yosef Weiss
Advised under Prof. Hagay Shpaisman and PhD student Udi Greenberg
During the summer of 2022, I was privileged to work in the Directed Materials Assembly laboratory in the Nanotechnology Department of Bar Ilan University. I joined a project exploring the use of laser-induced microbubbles for both two-dimensional and three-dimensional printing. The underlying mechanism requires focusing a laser beam at the interface between a substrate and a liquid containing nanoparticles or ions. Due to laser heating, microbubbles form and convection currents pin deposits at the liquid/bubble/substrate interface [1].
Figure 1: Laser-induced microbubble three-dimensional printing mechanism
Through the last nine years, this lab has developed a reliable method of two-dimensional, nanoparticle laser printing. Benefits of this printing format include achieving extremely narrow and precise lines (on the order of nanometers), the ability to print with a wide variety of nanoparticles, and a seamless transition between many different nanoparticle chemistries. With this technology, users can also stack multiple electrical connections by alternating between conductors and insulators.
When expanding this method to three-dimensional printing, two different techniques emerged—layer-by-layer printing and vector printing. Layer-by-layer printing builds up a design similar to a traditional three-dimensional printer; albeit using the laser technique. Vector printing directs the laser onto a continuously growing “stack” of material. For each of these printing styles, an issue arises. (1) For layer-by-layer printing, inconsistent deposition occurs due to sinusoidal patterns in the previous layer. The origin for this phenomenon is a self-enhanced process where areas with slightly more deposits tend to absorb more light, thus leading to even more deposits at the same location with each additional layer. (2) For vector printing, when the laser deviates from the vertical, pillar-like structure, deposits tend to accumulate unevenly at the bottom of the microbubble leading to deformation of the microstructure.
When approaching the former issue, the first step is to determine the optimal laser parameters for three-dimensional printing. Parametric properties include laser intensity, frequency, velocity, and duty cycle. Once two quality sets of parameters were identified, multiple layered lines were printed in order to visualize the pattern of deposition in the “wall” microstructure. Using a profilometer (optical probe), these layers were characterized in their height and roughness. It was previously determined that a shorter laser residence time results in less deposition due to a shorter period of nanoparticle heating [2]. To solve the problem of inconsistent deposition in layer-by-layer printing, a modulating laser was programmed to speed up and slow down depending on the height of the previous deposition. Through analyzing multiple examples, the period of the pattern of deposition was determined and then used for the modulation of the laser to correct (smooth) the deposition pattern.
For the latter issue, a double-axis stage was used to allow the nanoparticle solvent to be rotated to any orientation in all three dimensions. By rotating the stage, it keeps the “pillar” always parallel to gravity and therefore does not allow the structure to become unstable and irregular. A casing for these glass slides was designed and printed in-house in order to prevent the glass slide from falling off during rotations. A program was then written in order to calculate the degree of rotation for both axes and control the rotating stage to create the more complex shapes.
Developing the understanding and innovative tools for creating 3D structures with nanoparticles has a number of potentially significant applications. Some examples include creating compact stacked computer chips not possible with current technology, precision micro-motors and other mechanical devices, and micro-probes for visualization and measurement. The unique combination of micro/nano-size and high-precision will likely also lead to unanticipated novel applications.
References
- E. Greenberg et al., Adv. Mater. Interfaces, 1900541 (2019)
- N. Armon et al., ACS Appl. Mater. Interfaces, 9, 44214-44221 (2017)

Yedidya Moise
Yeshiva College, Yeshiva University, Class of 2024
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
Yonah Moise
Yeshiva College, Yeshiva University, Class of 2023
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Jeremy Schiff.
ABSTRACT:
This abstract has been previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book
Yonah Moise presented a poster on this research at the Yeshiva University’s 2023 STEM expo.
Yonah Moise also presented a poster on this research at Yeshiva University’s Katz Graduate School’s 2023 “Symposium on Science, Technology, and Health” An abstract authored by Yedidya Moise and Yonah Moise appeared in the symposium’s proceedings .
Title: Numerical Explorations in the Non-Linear Schrodinger Equation with Non-Symmetrical Gaussian Initial Conditions.
Keywords: Nonlinear Schrodinger equation, mathematical physics, applied math, computational math.

This research was conducted together with Yaelle Akhavan (SCW, ‘23) and Noa R. Atar (SCW, ‘22) – two more Yeshiva University students and participants of the YU/BIU Research Internship Program of Summer 2022.
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Eva Gilboa-Schechtman, PhD student Dan E. Hay, and Master’s students Noa Perets, Raquel Landau, and Itamar Zalkind.
These abstracts have been previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book .
Titles: "Emotional Congruence and Depression" and "Meta Analysis: Effects of Exclusion on Individuals with BPD."
Topic: Psychology.

SUMMARY:
This research focuses on understanding the properties and interactions of Codanin-1, a protein associated with Congenital Dyserythropoietic Anemia Type I (CDA I). The study investigates the influence of Codanin-1 on the activity and post-translational modification of Asf1, a histone chaperone protein, and explores methods for introducing a degron into the Codanin-1 gene to control protein degradation.
ABSTRACT:
Congenital Dyserythropoietic Anemia Type I (CDA 1); Exploring the Properties and Interactions of Codanin-1
Abigail Razi advised under Dr. Benny Motro and PhD student Moriya Kahta
Congenital Dyserythropoietic Anemia Type I (CDA I) is an autosomal recessive disease associated with abnormalities in erythrocyte precursor proliferation that may be expressed as spongy heterochromatin. The disorder is caused by a mutation in the CDANI gene encoding for the Codanin-1 protein. Codanin-1 is a conserved protein which serves as a protein scaffold and is presumably involved in chromatin assembly and maintenance.
Our research focuses on understanding the function of the Codanin-1 protein. Previously, it was demonstrated that Codanin-1 acts as a protein scaffold to recruit the histone chaperon protein Asf1. Overexpression of Codanin-1 results in a shift in the localization of Asf1 from the nuclear region to the cytoplasm.
In a subsequent analysis of the interactions between Codanin-1 and Asf1, the possible influence of Codanin-1 on post translational modification (PTM) of Asf1 was studied. To do so, HeLa cells underwent an inducible knockout of the gene that codes for Codanin-1. The prediction was that the knockout of this gene would prevent the expression of Codanin-1, thereby influencing the activity and PTM of Asf1. Then, immunoprecipitation was conducted to precipitate Asf1 from treated and untreated cells. Mass spectroscopy was used to identify PTM in Asf1 from cells containing endogenous Codanin-1 and cells containing the inducible knockout of Codanin-1. Differences in PTM between the endogenous Asf1 protein and the Asf1 protein following the inducible knockout of Codanin-1 would not only allude to the influence of Codanin-1 on the activity of Asf1, but more importantly, to larger revelations regarding the role of Codanin-1 in cellular function. The results of this study regarding Asf1 modifications have yet to be identified by mass spectroscopy.
In an additional project, I aimed to change the endogenous Codanin-1 gene by introducing a construct that contained a degron, just prior to its stop codon. A degron is a portion of a protein that is important for controlling protein degradation at the proteasome. In this study, the degron is Auxin (IAA)-dependent, namely in the presence of IAA receptors in the cells. When IAA is added, it binds to its receptor. The receptor recognizes the degron and sends Codanin-1 to the proteasome for degradation.
To insert the plasmid into the genome, a construct was created (referred to as #1217) by adding a 5’ arm and 3’ arm to allow for homologous recombination to occur. Crossover then occurred between the endogenous chromosome and the construct containing the degron and Hygromycin antibiotic resistance. The resulting chromosome contained the same DNA as the endogenous Codanin-1 gene, but also contained the additional degron and antibiotic resistance.
Because there are two chromosomes, it is necessary to introduce the degron into both strands using two constructs that confers cells resistant to both antibiotics: Hygromycin and Blasticidin. The introduction of this gene into the chromosome induces antibiotic resistance by producing a protein that degrades its respective antibiotic, thereby deeming the cell resistant. Since cells will only survive if antibiotic resistance has been introduced, cell survival would indicate successful transformation. In cells where the degron was inserted to both alleles, the addition of Auxin should lead to the degradation of Codanin-1.
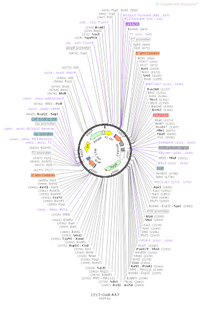
Using Crispr methodology, we utilized an sgRNA to cut a strand of chromosomal DNA in order to insert the degron which contained Hygromycin resistance. To insert the degron into the second chromosome, we looked to excise the Hygromycin antibiotic resistance fragment of the plasmid between NheI and NheI and replace it with Blasticidin antibiotic resistance. In order to insert the Blasticidin antibiotic resistant, we edited vector #1217, represented in Figure 1, by inserting a fragment from plasmid #1158 (Blasticidin resistance) in place of Hygromycin resistance. The construct conferring Blasticidin resistance will be introduced to the chromosome using Crispr methodology.
Figure 1. Gene map representation of vector #1217.
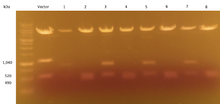
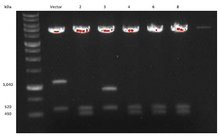
In order to examine if the transformation was successful, DNA from eight bacterial colony samples were run using gel electrophoresis. The results are demonstrated below in Figure 2. A band appearing around 1,040 kDa in samples Vector, 3, 5, and 7 is representative of the NheI-NheI fragment conferring Hygromycin resistance. This band indicates that these bacterial samples were not successfully transformed into containing the Blasticidin resistance. In all samples, a band present at 560 kDa is representative of the fragment between HindIII and HindIII. However, an additional band appeared for samples 2, 4, 6, and 8 around 490 kDa. Since it was difficult to distinguish between the band at 560 kDa and 490 kDa, we ran another gel with samples Vectors 2, 3, 4, 6, and 8. The results are demonstrated in Figure 3, where a band at 560 kDa is visible in all samples. Furthermore, there is a clear appearance of a band at 490 kDa in samples 2, 4, 6, and 8. Since the original NheI-NheI Hygromycin resistance sequence was 1,040 kDa long and was replaced with an NheI-NheI fragment for Blasticidin resistance that was 420 kDa long, the final sequence between ClaI and HindIII should appear around 490 kDa. The presence of this band identifies the samples which have been successfully transfected with Blasticidin resistance.
Figure 2. Transformation results examined using gel electrophoresis to compare vector #1217 and eight bacterial samples.
Figure 3. Transformation results repeated using gel electrophoresis. Bacterial samples 2, 4, 6, and 8 demonstrate successful transformation with Blasticidin resistance and, therefore, the degron.
The transformation described above is planned to be used in several mammalian cell types including HeLa, C12, U2OS, and SH-SY5Y cells. To study the effects of the transformation on Codanin-1, Auxin was first introduced into transformed HeLa cells. Interestingly, the knockout of Codanin-1 by the degron did not result in cell death or changes in the cell phenotype. This led to the theory that Codanin-1 is not required in all cell types. Thus, further studies in this area point to the knockout of Codanin-1 in cells in which Codanin-1 is believed to be of significance, such as U2OS or SHSY cells. The effects of Codanin-1 elimination on cellular function following introduction of Auxin in successfully transformed U2OS or SHSY cells has yet to be confirmed.
Previous studies we have conducted have worked towards determining the correct concentration of Hygromycin to use in order to achieve resistance. However, these studies have been unsuccessful as the concentration used have been found to be lethal. Further study of the optimal concentration of Hygromycin should be performed.
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Benny Motro and of PhD student Moriya Kahta.
This abstract was previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book.
Title: Congenital Dyserythropoietic Anemia Type I (CDA 1); Exploring the Properties and Interactions of Codanin-1.
Topic: Biology.

Washington University in St. Louis
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Hagay Shpaisman and of PhD student Udi Greenberg.
This abstract was previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book .
Title: Three-dimensional nanoparticle assembly by a modulated laser-induced microbubble for fabrication of a micrometric pattern.
Topic: Engineering, nanotechnology, directed materials assembly.
ABSTRACT:
Three-dimensional nanoparticle assembly by a modulated laser-induced microbubble for fabrication of a micrometric pattern
Yosef Weiss
Advised under Prof. Hagay Shpaisman and PhD student Udi Greenberg
During the summer of 2022, I was privileged to work in the Directed Materials Assembly laboratory in the Nanotechnology Department of Bar Ilan University. I joined a project exploring the use of laser-induced microbubbles for both two-dimensional and three-dimensional printing. The underlying mechanism requires focusing a laser beam at the interface between a substrate and a liquid containing nanoparticles or ions. Due to laser heating, microbubbles form and convection currents pin deposits at the liquid/bubble/substrate interface [1].
Figure 1: Laser-induced microbubble three-dimensional printing mechanism
Through the last nine years, this lab has developed a reliable method of two-dimensional, nanoparticle laser printing. Benefits of this printing format include achieving extremely narrow and precise lines (on the order of nanometers), the ability to print with a wide variety of nanoparticles, and a seamless transition between many different nanoparticle chemistries. With this technology, users can also stack multiple electrical connections by alternating between conductors and insulators.
When expanding this method to three-dimensional printing, two different techniques emerged—layer-by-layer printing and vector printing. Layer-by-layer printing builds up a design similar to a traditional three-dimensional printer; albeit using the laser technique. Vector printing directs the laser onto a continuously growing “stack” of material. For each of these printing styles, an issue arises. (1) For layer-by-layer printing, inconsistent deposition occurs due to sinusoidal patterns in the previous layer. The origin for this phenomenon is a self-enhanced process where areas with slightly more deposits tend to absorb more light, thus leading to even more deposits at the same location with each additional layer. (2) For vector printing, when the laser deviates from the vertical, pillar-like structure, deposits tend to accumulate unevenly at the bottom of the microbubble leading to deformation of the microstructure.
When approaching the former issue, the first step is to determine the optimal laser parameters for three-dimensional printing. Parametric properties include laser intensity, frequency, velocity, and duty cycle. Once two quality sets of parameters were identified, multiple layered lines were printed in order to visualize the pattern of deposition in the “wall” microstructure. Using a profilometer (optical probe), these layers were characterized in their height and roughness. It was previously determined that a shorter laser residence time results in less deposition due to a shorter period of nanoparticle heating [2]. To solve the problem of inconsistent deposition in layer-by-layer printing, a modulating laser was programmed to speed up and slow down depending on the height of the previous deposition. Through analyzing multiple examples, the period of the pattern of deposition was determined and then used for the modulation of the laser to correct (smooth) the deposition pattern.
For the latter issue, a double-axis stage was used to allow the nanoparticle solvent to be rotated to any orientation in all three dimensions. By rotating the stage, it keeps the “pillar” always parallel to gravity and therefore does not allow the structure to become unstable and irregular. A casing for these glass slides was designed and printed in-house in order to prevent the glass slide from falling off during rotations. A program was then written in order to calculate the degree of rotation for both axes and control the rotating stage to create the more complex shapes.
Developing the understanding and innovative tools for creating 3D structures with nanoparticles has a number of potentially significant applications. Some examples include creating compact stacked computer chips not possible with current technology, precision micro-motors and other mechanical devices, and micro-probes for visualization and measurement. The unique combination of micro/nano-size and high-precision will likely also lead to unanticipated novel applications.
References
- E. Greenberg et al., Adv. Mater. Interfaces, 1900541 (2019)
- N. Armon et al., ACS Appl. Mater. Interfaces, 9, 44214-44221 (2017)

Yedidya Moise
Yeshiva College, Yeshiva University, Class of 2024
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
Yonah Moise
Yeshiva College, Yeshiva University, Class of 2023
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Jeremy Schiff.
ABSTRACT:
This abstract has been previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book
Yonah Moise presented a poster on this research at the Yeshiva University’s 2023 STEM expo.
Yonah Moise also presented a poster on this research at Yeshiva University’s Katz Graduate School’s 2023 “Symposium on Science, Technology, and Health” An abstract authored by Yedidya Moise and Yonah Moise appeared in the symposium’s proceedings .
Title: Numerical Explorations in the Non-Linear Schrodinger Equation with Non-Symmetrical Gaussian Initial Conditions.
Keywords: Nonlinear Schrodinger equation, mathematical physics, applied math, computational math.

This research was conducted together with Yaelle Akhavan (SCW, ‘23) and Noa R. Atar (SCW, ‘22) – two more Yeshiva University students and participants of the YU/BIU Research Internship Program of Summer 2022.
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Eva Gilboa-Schechtman, PhD student Dan E. Hay, and Master’s students Noa Perets, Raquel Landau, and Itamar Zalkind.
These abstracts have been previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book .
Titles: "Emotional Congruence and Depression" and "Meta Analysis: Effects of Exclusion on Individuals with BPD."
Topic: Psychology.

SUMMARY:
This research focuses on understanding the properties and interactions of Codanin-1, a protein associated with Congenital Dyserythropoietic Anemia Type I (CDA I). The study investigates the influence of Codanin-1 on the activity and post-translational modification of Asf1, a histone chaperone protein, and explores methods for introducing a degron into the Codanin-1 gene to control protein degradation.
ABSTRACT:
Congenital Dyserythropoietic Anemia Type I (CDA 1); Exploring the Properties and Interactions of Codanin-1
Abigail Razi advised under Dr. Benny Motro and PhD student Moriya Kahta
Congenital Dyserythropoietic Anemia Type I (CDA I) is an autosomal recessive disease associated with abnormalities in erythrocyte precursor proliferation that may be expressed as spongy heterochromatin. The disorder is caused by a mutation in the CDANI gene encoding for the Codanin-1 protein. Codanin-1 is a conserved protein which serves as a protein scaffold and is presumably involved in chromatin assembly and maintenance.
Our research focuses on understanding the function of the Codanin-1 protein. Previously, it was demonstrated that Codanin-1 acts as a protein scaffold to recruit the histone chaperon protein Asf1. Overexpression of Codanin-1 results in a shift in the localization of Asf1 from the nuclear region to the cytoplasm.
In a subsequent analysis of the interactions between Codanin-1 and Asf1, the possible influence of Codanin-1 on post translational modification (PTM) of Asf1 was studied. To do so, HeLa cells underwent an inducible knockout of the gene that codes for Codanin-1. The prediction was that the knockout of this gene would prevent the expression of Codanin-1, thereby influencing the activity and PTM of Asf1. Then, immunoprecipitation was conducted to precipitate Asf1 from treated and untreated cells. Mass spectroscopy was used to identify PTM in Asf1 from cells containing endogenous Codanin-1 and cells containing the inducible knockout of Codanin-1. Differences in PTM between the endogenous Asf1 protein and the Asf1 protein following the inducible knockout of Codanin-1 would not only allude to the influence of Codanin-1 on the activity of Asf1, but more importantly, to larger revelations regarding the role of Codanin-1 in cellular function. The results of this study regarding Asf1 modifications have yet to be identified by mass spectroscopy.
In an additional project, I aimed to change the endogenous Codanin-1 gene by introducing a construct that contained a degron, just prior to its stop codon. A degron is a portion of a protein that is important for controlling protein degradation at the proteasome. In this study, the degron is Auxin (IAA)-dependent, namely in the presence of IAA receptors in the cells. When IAA is added, it binds to its receptor. The receptor recognizes the degron and sends Codanin-1 to the proteasome for degradation.
To insert the plasmid into the genome, a construct was created (referred to as #1217) by adding a 5’ arm and 3’ arm to allow for homologous recombination to occur. Crossover then occurred between the endogenous chromosome and the construct containing the degron and Hygromycin antibiotic resistance. The resulting chromosome contained the same DNA as the endogenous Codanin-1 gene, but also contained the additional degron and antibiotic resistance.
Because there are two chromosomes, it is necessary to introduce the degron into both strands using two constructs that confers cells resistant to both antibiotics: Hygromycin and Blasticidin. The introduction of this gene into the chromosome induces antibiotic resistance by producing a protein that degrades its respective antibiotic, thereby deeming the cell resistant. Since cells will only survive if antibiotic resistance has been introduced, cell survival would indicate successful transformation. In cells where the degron was inserted to both alleles, the addition of Auxin should lead to the degradation of Codanin-1.
Using Crispr methodology, we utilized an sgRNA to cut a strand of chromosomal DNA in order to insert the degron which contained Hygromycin resistance. To insert the degron into the second chromosome, we looked to excise the Hygromycin antibiotic resistance fragment of the plasmid between NheI and NheI and replace it with Blasticidin antibiotic resistance. In order to insert the Blasticidin antibiotic resistant, we edited vector #1217, represented in Figure 1, by inserting a fragment from plasmid #1158 (Blasticidin resistance) in place of Hygromycin resistance. The construct conferring Blasticidin resistance will be introduced to the chromosome using Crispr methodology.
Figure 1. Gene map representation of vector #1217.
In order to examine if the transformation was successful, DNA from eight bacterial colony samples were run using gel electrophoresis. The results are demonstrated below in Figure 2. A band appearing around 1,040 kDa in samples Vector, 3, 5, and 7 is representative of the NheI-NheI fragment conferring Hygromycin resistance. This band indicates that these bacterial samples were not successfully transformed into containing the Blasticidin resistance. In all samples, a band present at 560 kDa is representative of the fragment between HindIII and HindIII. However, an additional band appeared for samples 2, 4, 6, and 8 around 490 kDa. Since it was difficult to distinguish between the band at 560 kDa and 490 kDa, we ran another gel with samples Vectors 2, 3, 4, 6, and 8. The results are demonstrated in Figure 3, where a band at 560 kDa is visible in all samples. Furthermore, there is a clear appearance of a band at 490 kDa in samples 2, 4, 6, and 8. Since the original NheI-NheI Hygromycin resistance sequence was 1,040 kDa long and was replaced with an NheI-NheI fragment for Blasticidin resistance that was 420 kDa long, the final sequence between ClaI and HindIII should appear around 490 kDa. The presence of this band identifies the samples which have been successfully transfected with Blasticidin resistance.
Figure 2. Transformation results examined using gel electrophoresis to compare vector #1217 and eight bacterial samples.
Figure 3. Transformation results repeated using gel electrophoresis. Bacterial samples 2, 4, 6, and 8 demonstrate successful transformation with Blasticidin resistance and, therefore, the degron.
The transformation described above is planned to be used in several mammalian cell types including HeLa, C12, U2OS, and SH-SY5Y cells. To study the effects of the transformation on Codanin-1, Auxin was first introduced into transformed HeLa cells. Interestingly, the knockout of Codanin-1 by the degron did not result in cell death or changes in the cell phenotype. This led to the theory that Codanin-1 is not required in all cell types. Thus, further studies in this area point to the knockout of Codanin-1 in cells in which Codanin-1 is believed to be of significance, such as U2OS or SHSY cells. The effects of Codanin-1 elimination on cellular function following introduction of Auxin in successfully transformed U2OS or SHSY cells has yet to be confirmed.
Previous studies we have conducted have worked towards determining the correct concentration of Hygromycin to use in order to achieve resistance. However, these studies have been unsuccessful as the concentration used have been found to be lethal. Further study of the optimal concentration of Hygromycin should be performed.
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Benny Motro and of PhD student Moriya Kahta.
This abstract was previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book.
Title: Congenital Dyserythropoietic Anemia Type I (CDA 1); Exploring the Properties and Interactions of Codanin-1.
Topic: Biology.

Washington University in St. Louis
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Hagay Shpaisman and of PhD student Udi Greenberg.
This abstract was previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book .
Title: Three-dimensional nanoparticle assembly by a modulated laser-induced microbubble for fabrication of a micrometric pattern.
Topic: Engineering, nanotechnology, directed materials assembly.
ABSTRACT:
Three-dimensional nanoparticle assembly by a modulated laser-induced microbubble for fabrication of a micrometric pattern
Yosef Weiss
Advised under Prof. Hagay Shpaisman and PhD student Udi Greenberg
During the summer of 2022, I was privileged to work in the Directed Materials Assembly laboratory in the Nanotechnology Department of Bar Ilan University. I joined a project exploring the use of laser-induced microbubbles for both two-dimensional and three-dimensional printing. The underlying mechanism requires focusing a laser beam at the interface between a substrate and a liquid containing nanoparticles or ions. Due to laser heating, microbubbles form and convection currents pin deposits at the liquid/bubble/substrate interface [1].
Figure 1: Laser-induced microbubble three-dimensional printing mechanism
Through the last nine years, this lab has developed a reliable method of two-dimensional, nanoparticle laser printing. Benefits of this printing format include achieving extremely narrow and precise lines (on the order of nanometers), the ability to print with a wide variety of nanoparticles, and a seamless transition between many different nanoparticle chemistries. With this technology, users can also stack multiple electrical connections by alternating between conductors and insulators.
When expanding this method to three-dimensional printing, two different techniques emerged—layer-by-layer printing and vector printing. Layer-by-layer printing builds up a design similar to a traditional three-dimensional printer; albeit using the laser technique. Vector printing directs the laser onto a continuously growing “stack” of material. For each of these printing styles, an issue arises. (1) For layer-by-layer printing, inconsistent deposition occurs due to sinusoidal patterns in the previous layer. The origin for this phenomenon is a self-enhanced process where areas with slightly more deposits tend to absorb more light, thus leading to even more deposits at the same location with each additional layer. (2) For vector printing, when the laser deviates from the vertical, pillar-like structure, deposits tend to accumulate unevenly at the bottom of the microbubble leading to deformation of the microstructure.
When approaching the former issue, the first step is to determine the optimal laser parameters for three-dimensional printing. Parametric properties include laser intensity, frequency, velocity, and duty cycle. Once two quality sets of parameters were identified, multiple layered lines were printed in order to visualize the pattern of deposition in the “wall” microstructure. Using a profilometer (optical probe), these layers were characterized in their height and roughness. It was previously determined that a shorter laser residence time results in less deposition due to a shorter period of nanoparticle heating [2]. To solve the problem of inconsistent deposition in layer-by-layer printing, a modulating laser was programmed to speed up and slow down depending on the height of the previous deposition. Through analyzing multiple examples, the period of the pattern of deposition was determined and then used for the modulation of the laser to correct (smooth) the deposition pattern.
For the latter issue, a double-axis stage was used to allow the nanoparticle solvent to be rotated to any orientation in all three dimensions. By rotating the stage, it keeps the “pillar” always parallel to gravity and therefore does not allow the structure to become unstable and irregular. A casing for these glass slides was designed and printed in-house in order to prevent the glass slide from falling off during rotations. A program was then written in order to calculate the degree of rotation for both axes and control the rotating stage to create the more complex shapes.
Developing the understanding and innovative tools for creating 3D structures with nanoparticles has a number of potentially significant applications. Some examples include creating compact stacked computer chips not possible with current technology, precision micro-motors and other mechanical devices, and micro-probes for visualization and measurement. The unique combination of micro/nano-size and high-precision will likely also lead to unanticipated novel applications.
References
- E. Greenberg et al., Adv. Mater. Interfaces, 1900541 (2019)
- N. Armon et al., ACS Appl. Mater. Interfaces, 9, 44214-44221 (2017)

Yedidya Moise
Yeshiva College, Yeshiva University, Class of 2024
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
Yonah Moise
Yeshiva College, Yeshiva University, Class of 2023
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Jeremy Schiff.
ABSTRACT:
This abstract has been previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book
Yonah Moise presented a poster on this research at the Yeshiva University’s 2023 STEM expo.
Yonah Moise also presented a poster on this research at Yeshiva University’s Katz Graduate School’s 2023 “Symposium on Science, Technology, and Health” An abstract authored by Yedidya Moise and Yonah Moise appeared in the symposium’s proceedings .
Title: Numerical Explorations in the Non-Linear Schrodinger Equation with Non-Symmetrical Gaussian Initial Conditions.
Keywords: Nonlinear Schrodinger equation, mathematical physics, applied math, computational math.

This research was conducted together with Yaelle Akhavan (SCW, ‘23) and Noa R. Atar (SCW, ‘22) – two more Yeshiva University students and participants of the YU/BIU Research Internship Program of Summer 2022.
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Eva Gilboa-Schechtman, PhD student Dan E. Hay, and Master’s students Noa Perets, Raquel Landau, and Itamar Zalkind.
These abstracts have been previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book .
Titles: "Emotional Congruence and Depression" and "Meta Analysis: Effects of Exclusion on Individuals with BPD."
Topic: Psychology.

SUMMARY:
This research focuses on understanding the properties and interactions of Codanin-1, a protein associated with Congenital Dyserythropoietic Anemia Type I (CDA I). The study investigates the influence of Codanin-1 on the activity and post-translational modification of Asf1, a histone chaperone protein, and explores methods for introducing a degron into the Codanin-1 gene to control protein degradation.
ABSTRACT:
Congenital Dyserythropoietic Anemia Type I (CDA 1); Exploring the Properties and Interactions of Codanin-1
Abigail Razi advised under Dr. Benny Motro and PhD student Moriya Kahta
Congenital Dyserythropoietic Anemia Type I (CDA I) is an autosomal recessive disease associated with abnormalities in erythrocyte precursor proliferation that may be expressed as spongy heterochromatin. The disorder is caused by a mutation in the CDANI gene encoding for the Codanin-1 protein. Codanin-1 is a conserved protein which serves as a protein scaffold and is presumably involved in chromatin assembly and maintenance.
Our research focuses on understanding the function of the Codanin-1 protein. Previously, it was demonstrated that Codanin-1 acts as a protein scaffold to recruit the histone chaperon protein Asf1. Overexpression of Codanin-1 results in a shift in the localization of Asf1 from the nuclear region to the cytoplasm.
In a subsequent analysis of the interactions between Codanin-1 and Asf1, the possible influence of Codanin-1 on post translational modification (PTM) of Asf1 was studied. To do so, HeLa cells underwent an inducible knockout of the gene that codes for Codanin-1. The prediction was that the knockout of this gene would prevent the expression of Codanin-1, thereby influencing the activity and PTM of Asf1. Then, immunoprecipitation was conducted to precipitate Asf1 from treated and untreated cells. Mass spectroscopy was used to identify PTM in Asf1 from cells containing endogenous Codanin-1 and cells containing the inducible knockout of Codanin-1. Differences in PTM between the endogenous Asf1 protein and the Asf1 protein following the inducible knockout of Codanin-1 would not only allude to the influence of Codanin-1 on the activity of Asf1, but more importantly, to larger revelations regarding the role of Codanin-1 in cellular function. The results of this study regarding Asf1 modifications have yet to be identified by mass spectroscopy.
In an additional project, I aimed to change the endogenous Codanin-1 gene by introducing a construct that contained a degron, just prior to its stop codon. A degron is a portion of a protein that is important for controlling protein degradation at the proteasome. In this study, the degron is Auxin (IAA)-dependent, namely in the presence of IAA receptors in the cells. When IAA is added, it binds to its receptor. The receptor recognizes the degron and sends Codanin-1 to the proteasome for degradation.
To insert the plasmid into the genome, a construct was created (referred to as #1217) by adding a 5’ arm and 3’ arm to allow for homologous recombination to occur. Crossover then occurred between the endogenous chromosome and the construct containing the degron and Hygromycin antibiotic resistance. The resulting chromosome contained the same DNA as the endogenous Codanin-1 gene, but also contained the additional degron and antibiotic resistance.
Because there are two chromosomes, it is necessary to introduce the degron into both strands using two constructs that confers cells resistant to both antibiotics: Hygromycin and Blasticidin. The introduction of this gene into the chromosome induces antibiotic resistance by producing a protein that degrades its respective antibiotic, thereby deeming the cell resistant. Since cells will only survive if antibiotic resistance has been introduced, cell survival would indicate successful transformation. In cells where the degron was inserted to both alleles, the addition of Auxin should lead to the degradation of Codanin-1.
Using Crispr methodology, we utilized an sgRNA to cut a strand of chromosomal DNA in order to insert the degron which contained Hygromycin resistance. To insert the degron into the second chromosome, we looked to excise the Hygromycin antibiotic resistance fragment of the plasmid between NheI and NheI and replace it with Blasticidin antibiotic resistance. In order to insert the Blasticidin antibiotic resistant, we edited vector #1217, represented in Figure 1, by inserting a fragment from plasmid #1158 (Blasticidin resistance) in place of Hygromycin resistance. The construct conferring Blasticidin resistance will be introduced to the chromosome using Crispr methodology.
Figure 1. Gene map representation of vector #1217.
In order to examine if the transformation was successful, DNA from eight bacterial colony samples were run using gel electrophoresis. The results are demonstrated below in Figure 2. A band appearing around 1,040 kDa in samples Vector, 3, 5, and 7 is representative of the NheI-NheI fragment conferring Hygromycin resistance. This band indicates that these bacterial samples were not successfully transformed into containing the Blasticidin resistance. In all samples, a band present at 560 kDa is representative of the fragment between HindIII and HindIII. However, an additional band appeared for samples 2, 4, 6, and 8 around 490 kDa. Since it was difficult to distinguish between the band at 560 kDa and 490 kDa, we ran another gel with samples Vectors 2, 3, 4, 6, and 8. The results are demonstrated in Figure 3, where a band at 560 kDa is visible in all samples. Furthermore, there is a clear appearance of a band at 490 kDa in samples 2, 4, 6, and 8. Since the original NheI-NheI Hygromycin resistance sequence was 1,040 kDa long and was replaced with an NheI-NheI fragment for Blasticidin resistance that was 420 kDa long, the final sequence between ClaI and HindIII should appear around 490 kDa. The presence of this band identifies the samples which have been successfully transfected with Blasticidin resistance.
Figure 2. Transformation results examined using gel electrophoresis to compare vector #1217 and eight bacterial samples.
Figure 3. Transformation results repeated using gel electrophoresis. Bacterial samples 2, 4, 6, and 8 demonstrate successful transformation with Blasticidin resistance and, therefore, the degron.
The transformation described above is planned to be used in several mammalian cell types including HeLa, C12, U2OS, and SH-SY5Y cells. To study the effects of the transformation on Codanin-1, Auxin was first introduced into transformed HeLa cells. Interestingly, the knockout of Codanin-1 by the degron did not result in cell death or changes in the cell phenotype. This led to the theory that Codanin-1 is not required in all cell types. Thus, further studies in this area point to the knockout of Codanin-1 in cells in which Codanin-1 is believed to be of significance, such as U2OS or SHSY cells. The effects of Codanin-1 elimination on cellular function following introduction of Auxin in successfully transformed U2OS or SHSY cells has yet to be confirmed.
Previous studies we have conducted have worked towards determining the correct concentration of Hygromycin to use in order to achieve resistance. However, these studies have been unsuccessful as the concentration used have been found to be lethal. Further study of the optimal concentration of Hygromycin should be performed.
Yeshiva University Bar-Ilan University Summer Science Research Internship Program, Summer 2022
This research was conducted at Bar-Ilan University (Ramat-Gan, Israel) during Summer 2022, under the mentorship of Professor Benny Motro and of PhD student Moriya Kahta.
This abstract was previously published in the Yeshiva University Bar-Ilan University Summer Science Research Internship Program’s Summer 2022 abstract book.
Title: Congenital Dyserythropoietic Anemia Type I (CDA 1); Exploring the Properties and Interactions of Codanin-1.
Topic: Biology.